SIMON Says…
Researchers at Oxford Vaccine Group have developed machine learning software to predict the efficacy of flu vaccines, offering huge potential for other vaccine research.
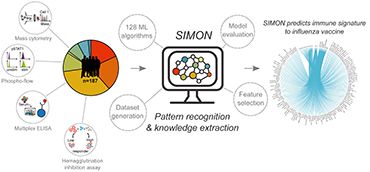 TOMIC – SIMON Says
TOMIC – SIMON Says(Credit: Dr Adriana Tomic)
Researchers at the University of Oxford Vaccine Group wanted to explore whether they could improve the effectiveness of the vaccine by identifying which individuals are likely to respond positively to receiving it.
As a first step, the team spent three years collaborating with researchers at Stanford University to develop software which can quickly analyse clinical data from multiple different studies. They then used ‘SIMON’ (Sequential Iterative Modeling OverNight) to analyse five studies conducted on the same flu vaccine between 2007-15 under the FluPRINT project.
“Data had been generated from different trials and were recorded in hundreds of different files,” explains Tomic. “Before we could use SIMON, we needed to pre-process, merge, clean, and standardise the data – which was very time consuming.”
“But it was worth it,” she continues. “When we were able to apply the machine learning process, we quickly identified subsets of immune cells that indicate that an individual with those cells would respond well to the vaccine. This could be hugely important in developing future vaccines as it may be possible to increase the number of those cell types in poor responders – and so improve the effectiveness of the vaccine for them.”
SIMON offers huge potential to transform the speed and efficacy of vaccine development well beyond the flu vaccine. Tomic says: “Machine learning can be extremely helpful in understanding biological processes because it enables rapid examination of huge amounts of data from different datasets – but unfortunately many biologists don’t have the computational skills to use it. SIMON can help them build thousands of models to analyse complex data, using more than 200 state-of-the-art algorithms, at the click of a button.”
SIMON is now used by leading experts in human immunology and vaccinology at the universities of Oxford and Stanford and the software is available free to download. “Many innovations in machine learning are restricted to users who can pay a high price for them,” says Tomic. “We believe that the software is not a commodity and should be freely available to all researchers.”
The team is currently applying SIMON to examine whether the protection offered by the flu vaccine in children is lowered by re-immunisation, which would profoundly change how the vaccine is administered. SIMON has also been successfully applied to other vaccines being tested by the Oxford Group, including meningococcus and salmonella typhi.
This work has already achieved significant impact. The first paper on SIMON was one of the 20 ‘most read manuscripts’ in 2019 according to the American Association of Immunologists and was selected as a finalist for the Award of Excellence from the International Society for Cytometry. The SIMON software has been downloaded 3000 times and could reach more than 100,000 potential users. The project website provides resources in multiple languages including English, Chinese, Arabic, French, and German.
Tomic is part of the University of Oxford team that has identified a potential vaccine for COVID-19 and is working toward the first clinical testing phase. She comments: “As understanding of the immune system improves and more data is collected, SIMON has huge potential to support data-driven research in vaccine development. Its application of machine learning to immunological data may also be helpful in the development of the COVID vaccine – and will be vital to developing more targeted and effective therapies in future.”
Dr Adriana Tomic is Marie Curie Fellow in the department of Paediatrics and a member of Oxford Vaccine Group.
Funders: EC Horizon2020.