Oxford-led study shows how AI can detect antibiotic resistance in as little as 30 minutes
To mark World Antimicrobial Awareness Week, researchers supported by the Oxford Martin Programme on Antimicrobial Resistance Testing at the University of Oxford have reported advances towards a novel and rapid antimicrobial susceptibility test that can return results within as little as 30 minutes - significantly faster than current gold-standard approaches.
In an era where AMR poses a critical public health threat, our team has made a ground-breaking advancement toward rapid detection of AMR. This innovation may hold the potential to revolutionise the way we respond to infectious diseases, allowing for more precise and timely treatment decisions, ultimately saving lives.
Co-author Dr Piers Turner, Oxford Martin Programme on Antimicrobial Resistance Testing and Department of Physics, University of Oxford
In their study published in Communications Biology, the team used a combination of fluorescence microscopy and artificial intelligence (AI) to detect antimicrobial resistance (AMR). This method relies on training deep-learning models to analyse bacterial cell images and detect structural changes that may occur in cells when they are treated with antibiotics. The method was shown to be effective across multiple antibiotics, achieving at least 80% accuracy on a per-cell basis.
The researchers say their model could be used to identify whether cells in clinical samples are resistant to a range of a wide variety of antibiotics in the future.
Co-author of the paper Achillefs Kapanidis, Professor of Biological Physics and Director of the Oxford Martin Programme on Antimicrobial Resistance Testing, said: ‘Antibiotics that stop the growth of bacterial cells also change how cells look under a microscope, and affect cellular structures such as the bacterial chromosome. Our AI-based approach detects such changes reliably and rapidly. Equally, if a cell is resistant, the changes we selected are absent, and this forms the basis for detecting antibiotic resistance.’
The researchers tested their method on a range of clinical isolates of E. coli, each with varying levels of resistance to the antibiotic ciprofloxacin. The deep-learning models were able to detect antibiotic resistance reliably and at least 10 times faster than established state-of-the art clinical methods considered to be gold standard.
The team hopes to continue developing their method so that it becomes faster and more scalable for clinical use, as well as adapting its usage for different types of bacteria and antibiotics.
According to the Global Research on Antimicrobial Resistance (GRAM) Project – a partnership involving the University – almost 1.3 million people died in 2019 due to AMR.
Current testing methods rely on growing bacterial colonies in the presence of antibiotics. However, such tests are slow, often requiring several days to understand how resistant bacteria are to a range of antibiotics.
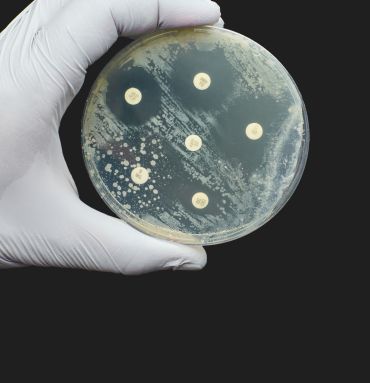 Current testing methods for antibiotic susceptibility rely on growing bacterial colonies in the presence of antibiotics – typically requiring several days. Image credit: Nicolae Malancea, Getty Images.
Current testing methods for antibiotic susceptibility rely on growing bacterial colonies in the presence of antibiotics – typically requiring several days. Image credit: Nicolae Malancea, Getty Images.The researchers say that if developed further, the rapid nature of their method may facilitate targeted antibiotic treatments – helping to decrease treatment times, minimise side effects, and ultimately slow down the rise of AMR.
Co-author of the paper Aleksander Zagajewski, doctoral student with the University’s Department of Physics, said: ‘Time is beginning to run out for our antibiotic arsenal; we are hoping our novel diagnostics will pave the way for a new generation of precision treatments for the most sick patients.’
The study ‘Deep learning and single-cell phenotyping for rapid antimicrobial susceptibility detection in Escherichia coli’ has been published in Communications Biology.
Researchers from across the University contributed to the study, including from the Department of Physics, Nuffield Department of Medicine, Sir William Dunn School of Pathology and Nuffield Department of Women’s Reproductive Health. Researchers from the Oxford University Hospitals NHS Foundation Trust’s Department of Microbiology and Infectious Diseases were also involved.
 Largest ever UK study reveals stark ethnic and social inequalities in lung cancer diagnosis
Largest ever UK study reveals stark ethnic and social inequalities in lung cancer diagnosis
 Oxford’s gargoyles come to life in new Extended Reality (XR) interactive experience
Oxford’s gargoyles come to life in new Extended Reality (XR) interactive experience
 Treating bullying as everyone’s problem reduces incidence in primary schools
Treating bullying as everyone’s problem reduces incidence in primary schools
 Oxford's Ashmolean Museum saves Fra Angelico masterpiece to go on public display from December
Oxford's Ashmolean Museum saves Fra Angelico masterpiece to go on public display from December
 New, ARIA-backed project aims to unlock radically cheaper AI hardware
New, ARIA-backed project aims to unlock radically cheaper AI hardware