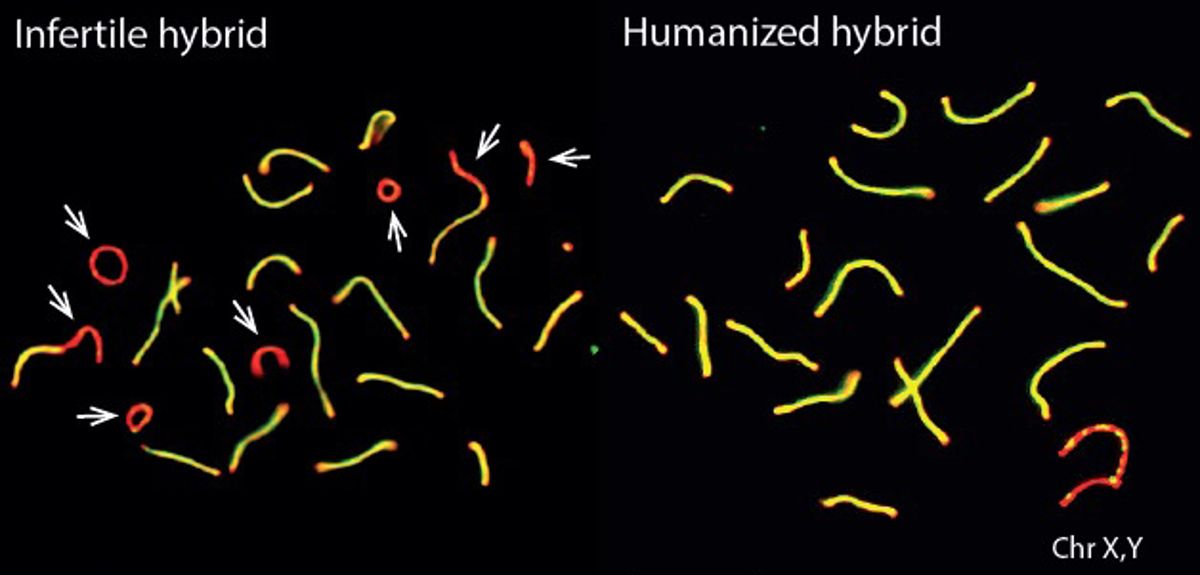
Image credit: Daniela Moralli
The origin of a species
A study by researchers at the Wellcome Trust Centre for Human Genetics at Oxford University has uncovered the key role played by a single gene in how groups of animals diverge to form new species. The study, published today in the journal Nature, restored fertility to the normally-infertile offspring of two subspecies of mice, by replacing part of the Prdm9 gene with the equivalent human version. Despite the nearly 150 million years of evolution separating mice and humans, these 'humanized' mice were completely fertile.
New animal species form when groups of animals become isolated and as a result, begin to separate through evolution (a process known as speciation). When these isolated populations meet later, they might be able to breed with each other, but the male offspring are often infertile. Horses and donkeys are an example of such speciation: they can interbreed, but their offspring, mules, are infertile.
'Our work studied similar infertility in hybrid house mice, whose two parents come from different subspecies found in Western and Eastern Europe', says Dr Ben Davies from the Nuffield Department of Medicine, the first author on the study. These two sub-species are therefore on the verge of splitting into two entirely different species, since like mules, their offspring are infertile.
Dr Davies and his colleagues studied the Prdm9 gene: this gene is already known to have a role in infertility in mice from different species, and is in fact the only known speciation gene in mammals. However, how speciation might link up to infertility was unknown.
A previous clue came from earlier work by Professor Simon Myers and Professor Peter Donnelly, the two senior authors on this study. Their work had found that the protein produced by the Prdm9 gene determines where in the genome maternal and paternal chromosomes exchange genetic information: a process known as recombination, which controls how genes are passed down through a species.
To figure out the exact involvement of the gene in how species form, the transgenic, chromosome dynamics and the genomics core facilities at the Wellcome Trust Centre for Human Genetics came together to carry out a new study. The groups replaced the region of the mouse Prdm9 gene responsible for DNA binding with the equivalent sequence from humans, thus completely changing where recombination happened along the genome.
'The effect of this change was startling', says Professor Myers. 'When these humanized mice were crossed with mice from the other subspecies, the offspring were no longer infertile but were instead fully fertile: inserting a key part of the human version of the gene into the mouse DNA binding domain had completely reversed the infertility of hybrid mice.' Speciation thus seems to be a potentially reversible event, at least in mice, where the human allele mimics what may happen when a random Prdm9 mutation occurs.
Only the binding properties of PRDM9 protein were changed in the experiment, so the researchers also investigated PRDM9 binding to DNA in more detail. They found that the infertile mouse hybrids showed a striking pattern: the mouse PRDM9 protein would bind to one of their chromosomes or the other, but not both. This happened even though the two copies of each chromosome they carry – one from each subspecies – are more than 98% similar overall.
The researchers discovered that this strange binding pattern came about because over many generations, the normal mouse PRDM9 protein erodes the DNA sequence it binds to, resulting in the asymmetric chromosome binding pattern seen in the infertile hybrids. The researchers think that asymmetric binding makes it more difficult for chromosomes to successfully identify and make contact with each other as egg and sperm cells are formed. The result is that in many different hybrid mice, asymmetric PRDM9 binding is associated with an increasing failure rate in chromosomes making contact correctly, leading to more and more fertility problems.
'We think that the symmetric marking of chromosomes by PRDM9 facilitates their pairing: where PRDM9 binding is very asymmetric, this leads to difficulties in pairing, failure in recombination repair and, at one extreme, the infertility we see in some mouse hybrids', says Professor Donnelly. 'These results also highlight just how important it is to understand the co-evolution of the Prdm9 gene with the whole genome in which it resides: we now have a new mechanism for reproductive isolation of closely related subspecies.'
The research team are now busy re-engineering different version of the Prdm9 gene to explore further, and they hope to find out how exactly the symmetry of DNA binding can influence the pairing of chromosomes.