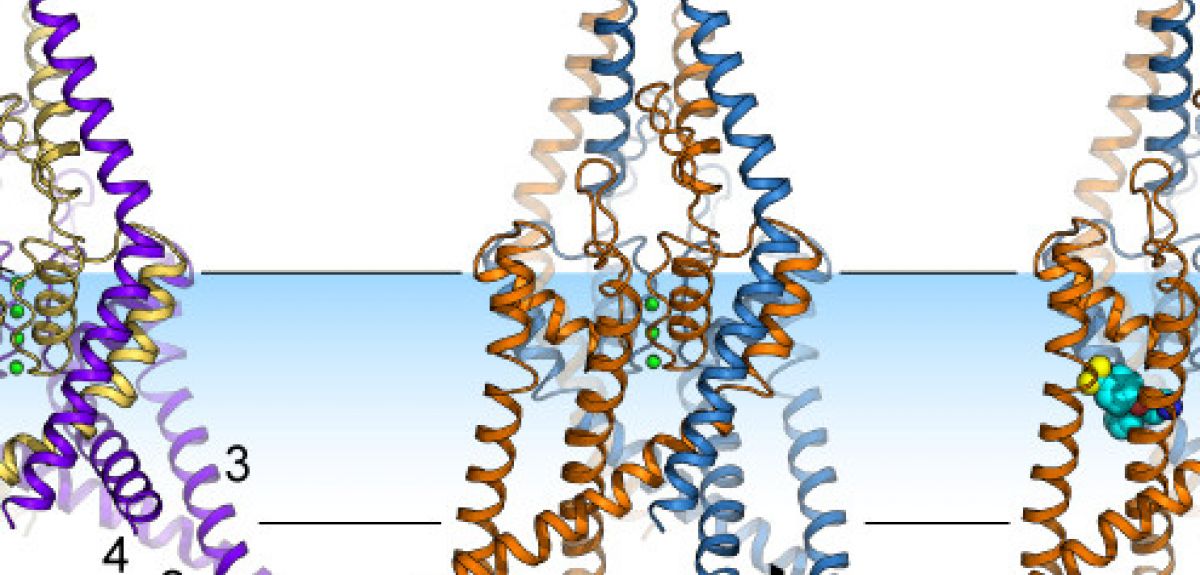
Image courtesy Liz Carpenter & Stephen Tucker
Hunting the Ion Channel
The electrical impulse that powers the workings of the brain and the heart begins with charged particles passing through cellular structures known as ion channels. Using the same technique used to decode the structure of DNA, an Oxford University team has been able to catch snapshots of an ion channel in action. Their results, published in Science earlier this month, help explain how neurons respond to a variety of stimuli, from drugs to touch.
I talked to the lead authors, Professor Liz Carpenter from the Structural Genomics Consortium and Professor Stephen Tucker from the Department of Physics, about why it Oxford is a particularly good place to study ion channels.
OxSciBlog: What are ion channels and why are they important?
Stephen Tucker: Ion channels are literally holes in the cell membrane, and they allow electrically charged particles ('ions') to move from one side of the cell membrane to the other. This is the process responsible for conducting the electrical signal via which the brain and the heart work.
So ion channels are important because they control neuron excitability, which in turn controls how different parts of our bodies talk to each other and how we sense the external world.
Liz Carpenter: Cell membranes are what separate the inside of cells from the outside, and ion channels sit right in the cell membrane. When a nerve fires a burst of electricity, charged particles flow through ion channels, acting like a current. This current then triggers other channel opening, and this is how electrical activity flows down the nerve cell.
OSB: You've got many collaborators from pharmaceutical companies as well: why are these cellular structures of particular interest for people interested in drug development?ST: They're many, many different kinds of ion channels, letting in different kinds of charged particles. We're specifically looking at an ion channel called TREK-2 that let in charged potassium ions, and while you find these channels all over the heart and the brain, they're especially concentrated in nerves that sense pain.
And if we can figure out a way to activate these channels, to keep them open, then we can actually silence the cascade of electrical activity through which neurons communicate. This could potentially block the pain signal from getting to the brain.
OSB: So how are these channels activated?
LC: Originally, people thought that these channels just let in charged potassium ions. But since then, researchers have found that they are activated by stretching the cell membrane, by temperature changes, and by a whole range of pharmaceutical compounds. We still don't know how all these ways of modifying the activity of these channels affects the activity of cells in the brain. What we do know is that in a test-tube, you add some drugs to these channels and they open, and other drugs make them close.
The problem was that before our study, the picture that we had for these ion channels was a little square blob: we didn't actually know what their structure looks like, or how they open and close.
So my group has been studying all possible human ion channels (they are 240 of them!). These proteins are very difficult to crystallize (which you need to do in order to study their structure), but the family of ion channels that TREK-2 belongs to are particularly well-behaved. Not only did we manage to crystallize the TREK-2 channel, but just by chance, we actually got crystals of two different states of the channel: we saw a structure that was stretched, and then another structure that was much less stretched.
OSB: What does the structure of these TREK-2 channels look like?
LC: This channel has two protein chains, with each chain made up of a few thousand atoms. Each chain folds up into a series of helices and loops, which come together to form a folded structure with a pore at the centre. This pore acts as a filter: the oxygen atoms in the pore are lined up in such a way that they can only bind potassium ions, so that is the only thing that can pass through these channels.
When we looked at the crystal structure of these ion channels, we found that the helices could either move down into the inside of the cell, or move up, into the cell membrane. This movement is somehow controlling the flow of ions past the oxygen atoms and through these channels.
OSB: How did you discover this structure?
LC: We have to start by persuading a 'reporter' line of insect cells to produce the protein that we're interested in studying: we splice the gene for the specific protein that we're interested in into an insect virus, which then infects the cells, causing them to make large amounts of the protein.
The next problem we have is that these ion channel proteins are water-repellent (hydrophobic) where they interact with the cell membrane, and water-attracting (hydrophilic) where the interact with the inside and outside of the cells. So we can't just pull these ion channels out of their cell membranes by dissolving them in water.
So we add detergent, which is good at dissolving hydrophobic and hydrophilic molecules: it's not quite washing up liquid that we’re using, but it's not far off!
Then we try use a variety of different reagents to try and persuade all these ion channels to line up in orderly arrays next to each other: to make crystals, in other words. Whether this works or not is very much a matter of luck, I'm afraid!
OSB: Why do you need to crystallize these molecules in order to study them?
LC: We work out the structure of molecules from the diffraction pattern that are produced by passing a beam of X-rays (at the Diamond light source in Harwell) through the material: single molecules by themselves aren't big enough to produce a strong enough diffraction pattern that we can study.
But if we line up each of the molecules to make a crystal, the crystal acts as an amplifier, producing a clear diffraction pattern. We rotated the crystal within the X-ray beam to get different diffraction patterns, and then we worked back from the diffraction patterns to figure out where each individual atom sits in the ion channel.
This is the same technique that was used to work out the structure of DNA, another complex protein molecule. But we're finally getting to the point that we can use this method to look at the structure of human proteins, which have been particularly hard to work with: they’re hard to express in reporter cell lines (if we get half a milligram of protein, we’re very happy!), and hard to crystallize.
OSB: What was the next step, once you had the structure worked out?
ST: We were lucky enough to solve more than one protein structure. But we still didn't know which structure corresponded to the open version of the channel and which to the closed version: just looking at two structures, you can't tell which one was open and which was closed, since it looked like ions could get through both forms of the channel.
So we took advantage of the fact that some drugs, such as Prozac, are known to only bind to the TREK-2 channel when it is closed. So when Liz managed to get a crystal of TREK-2 with Prozac bound to it, we knew that this version of TREK-2 had its 'pore' closed, not allowing potassium to travel through it.
LC: We also wanted to track exactly where Prozac binds to the channel. But Prozac itself doesn't show up in the diffraction patterns that we're using, so we used another trick: we attached a molecule of Bromine to the Prozac. Bromine lights up nicely in the diffraction pattern, and lets us see exactly where the Prozac is attaching itself to TREK-2.
So we really needed all sorts of different expertise in order to these experiments: my experience is in decoding protein structures, and Stephen's interest is ion channel function. But we needed the help of our colleagues from the Chemistry Department to do the Bromine tagging experiment, and colleagues at a pharmaceutical company supplied us with the drugs that we used in the experiment.
ST: In Oxford, we're lucky enough to have a huge consortium of people who work on ion channels (the OXION consortium), and I think this study demonstrates why Oxford has become a major international centre for this kind of work. For example, the two images that we got were frozen snapshots, but we think that this channel is likely to exist in many states, rather than just two. Computational modelling from other colleagues (Professor Mark Sansom in the Department of Biochemistry) was also essential to give us an idea of how the protein might actually move in a cell membrane.
OSB: What sort of future work are you planning?
ST: The drug that we looked at, Prozac, binds to a 'window' in the channel that only exists in its closed state. We'd like to look at how some other drugs bind to TREK-2, and we need very high-resolution 'pictures' of the channel in order to do this.
LC: There are other ion channels related to TREK-2 that we'd like to look at: one of them, TRESK, has been linked to migraines for example. We'd like to work out the structure of these channels, especially since many of them have been linked genetically to diseases. We're also trying to get more uniform crystals to increase the resolution of the structure we get.
We're also going to use a fantastic new device called the LINAC Coherent Light Source (LCLS) in Stanford: it is about a billion times more powerful that the light source we've used previously, which means that laser pulses hit the sample hundreds of times every second. What we're going to do here is fire a slurry of small crystals through the LCLS beam. The crystals explode as they hit the beam, but we get our data before the atoms within the crystal have time to move apart. The data we get from the LCLS is going to much higher-resolution, and it will allow us to study ion channels that don’t crystallize into large crystals.
A report of the research, entitled 'K2P channel gating mechanisms revealed by structures of TREK-2 and a complex with Prozac', is published in Science.
 Recreating stellar conditions on Earth
Recreating stellar conditions on Earth Lizards quickly adapt to chilly English climate
Lizards quickly adapt to chilly English climate Making sense of sex: why genes recombine
Making sense of sex: why genes recombine 8 things about Oxford’s driverless tech
8 things about Oxford’s driverless tech A song of fire and ice in the ocean
A song of fire and ice in the ocean