
Image credit: Shutterstock
Using portable, real time DNA sequencing to fight drug-resistant TB
Scientists from the Madagascar National TB Program, Institute Pasteur Madagascar, University of Oxford, European Bioinformatics Institute (EMBL-EBI) and Stony Brook University are collaborating to train Malagasy scientists to rapidly detect tuberculosis (TB) and drug-resistance using DNA sequencing. The goal of the project is to improve diagnosis and treatment, and will provide insights on disease transmission.
Scientists in Madagascar have for the first time performed DNA sequencing in-country using portable technology to rapidly identify the bacteria responsible for tuberculosis (TB) and its drug resistance profile. The project, led by an ambitious, global team of doctors and scientists from Madagascar’s National TB Program, Institute Pasteur Madagascar (IPM), University of Oxford, European Bioinformatics Institute (EMBL-EBI) and Stony Brook University, is seeking to transform the surveillance, diagnosis and treatment of TB and other infectious diseases in Madagascar. The researchers used the portable, affordable MinION sequencing device, developed by Oxford spin-out company Oxford Nanopore Technologies.
Beyond performing DNA sequencing on samples submitted to the nation reference laboratory for TB, the team partnered with TB clinics in the country to evaluate the ability to perform these analyses outside the labs or “in the field”. To achieve public health impact, their objective is to bring TB DNA sequencing closer to the patients for rapid diagnosis.
Niaina Rakotosamimanana, head of the mycobacterium unit at IPM said: 'This exercise has provided evidence that we can identify TB and its drug resistance properties using real time DNA sequencing technology. We have done this in the lab and are doing more development towards providing this in a rural setting without traditional lab facilities and very limited resources. This is exactly where technology is needed – close to where the TB outbreaks are actually taking place.'
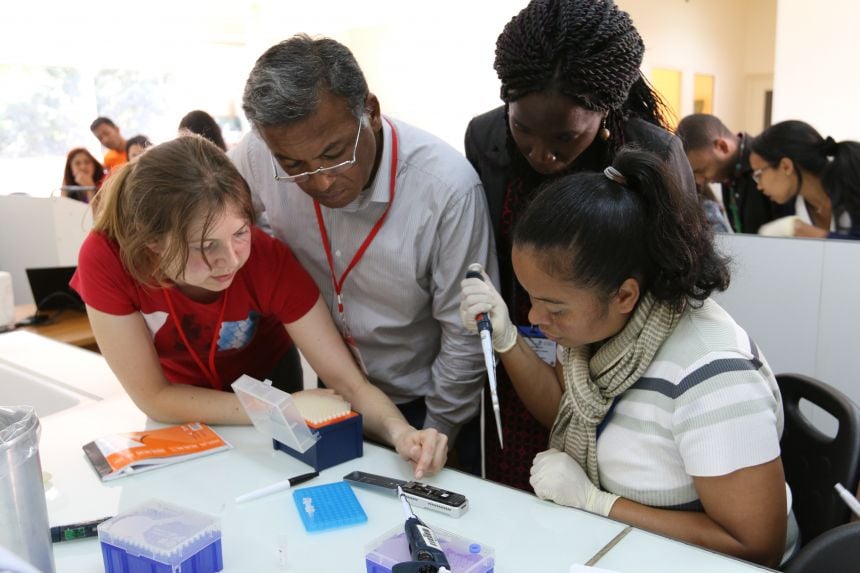 Using portable, real time DNA sequencing to fight drug-resistant TB
Using portable, real time DNA sequencing to fight drug-resistant TBImage credit: Zoe McDougall (Oxford Nanopore Technologies)
Simon Grandjean Lapierre, Research Coordinator at Stony Brook University said: 'We have trained 22 scientists to perform DNA sequencing using the MinION platform. This will allow better characterization and understanding of a variety of infectious diseases which represent public health threats in Madagascar. Our long term goal, as part of our broader TB strategy, is to increase capacity and work on methods development so that DNA sequencing can become part of routine TB diagnosis and surveillance in Madagascar.'
In high-income countries, DNA sequencing of TB samples is increasingly being deployed for diagnosis and surveillance of the disease. It delivers rich information about potential drug resistant bacteria, and can identify transmission between patients, with a level of precision and speed unachievable by the traditional methods. However, due to the cost and complexity of DNA sequencing systems, these insights have so far remained out of reach of low- and middle-income countries, where TB has the greatest health impact. The MinION increases DNA sequencing technology accessibility, due to its portability and cost.
Genomic surveillance provides opportunities to gain rich information about the pathogens that are affecting a population. The impact of access to DNA sequencing in Madagascar will not be limited to tackling TB. The scientists from Institut Pasteur de Madagascar have also now sequenced a range of viruses and bacteria that are endemic to the country. Complete genomes of pathogens including Coronavirus (a variant of SARS and MERS-CoV), Respiratory Syncytial Virus (the common cold virus), carbapenemase-producing strains of Klebsiella pneumonia (a bacteria highly resistant to antibiotics) and Yersinia pestis (the bacteria responsible for plague) have been sequenced. Insights from these studies could help to bring future outbreaks of these diseases under control more quickly.
The training led by the Institut Pasteur Madagascar, Oxford University, EMBL-EBI and Stony Brook University, will be extended to healthcare facilities of the National TB Program of Madagascar.
 Using portable, real time DNA sequencing to fight drug-resistant TB
Using portable, real time DNA sequencing to fight drug-resistant TBImage credit: Zoe McDougall (Oxford Nanopore Technologies)
As part of a larger study funded by the Wellcome Trust, the Academy of Medical Sciences and NIHR Oxford Biomedical Research Centre the team of researchers aim to develop more capacity for DNA sequencing and use this tool prospectively to identify TB drug resistance, better understand the dynamics of TB transmission in Madagascar and evaluate impact on public health.
 Oxford tops QS World University Rankings in four subjects, named overall top for Humanities
Oxford tops QS World University Rankings in four subjects, named overall top for Humanities
 Blood test may improve survival of childhood cancer in Africa
Blood test may improve survival of childhood cancer in Africa