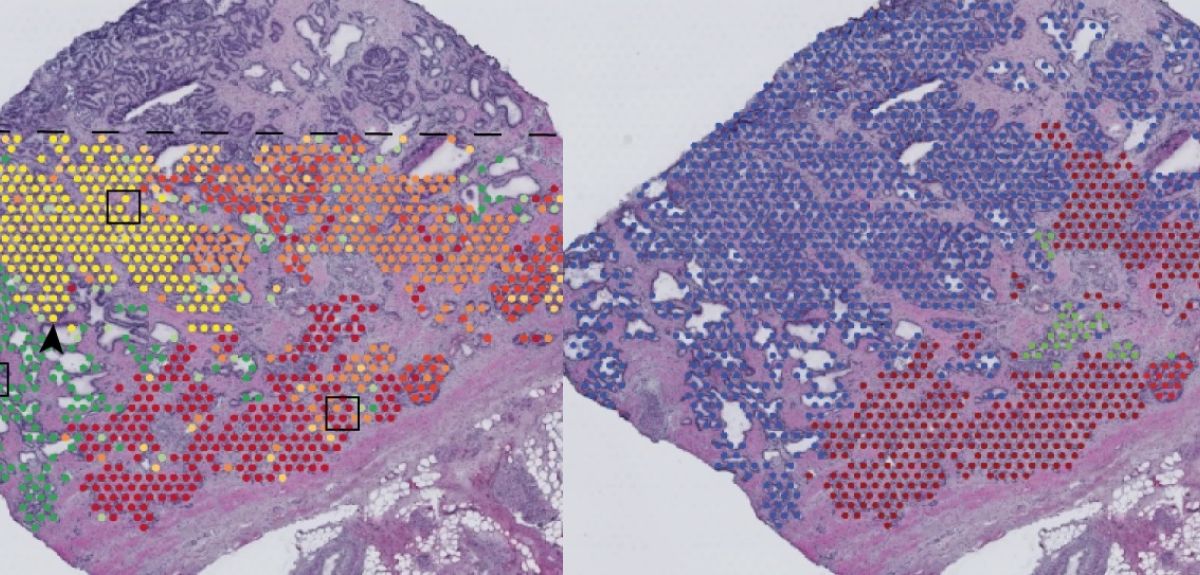
Image credit: Nature
Genetic mapping of tumours reveals how cancers grow
Researchers from the University of Oxford, KTH Royal Institute of Technology, Science for Life Laboratory, and the Karolinska Institutet, Solna, Sweden, have found that individual prostate tumours contain a previously unknown range of genetic variation.
Understanding which cells give rise to which areas of cancer can improve our understanding of how a tumour has grown and developed, including how it has changed genetically, over time. This has been made possible using a new technique called spatial transcriptomics, which allows scientists to see what genetic changes take place without breaking up the tissue they’re looking at. This adds a new dimension which researchers have now used to reveal which cells have mutated and where within the ecosystem of an organ.
Current techniques for studying the genetics of cells within tumours involve taking a sample from the cancerous area and analysing the DNA of those cells. The problem is that many cancers, such as prostate cancer, are three dimensional, which means that any one sample would only give a small snapshot of the tumour.
In a new study published in Nature, the researchers used spatial transcriptomics to create a cross-sectional map of a whole prostate, including areas of healthy and cancerous cells. By grouping cells according to similar genetic identity, they were surprised to see areas of supposedly healthy tissue that already had many of the genetic characteristics of cancer. This finding was surprising because of both the genetic variability within the tissue as well as the large number of cells that would be considered healthy, but which contained mutations usually identified with cancerous cells.
Alastair Lamb of Oxford’s Nuffield Department of Surgical Sciences, who jointly led the study, said: 'Prostate tissue is three-dimensional, and like most organs that can develop cancer we still have much to learn about what cellular changes cause cancer and where it starts. One thing we are fairly confident of is that it starts with genetic mutations.
'We have never had this level of resolution available before, and this new approach revealed some surprising results. For example, we have found that many of the copy number events we previously thought to be linked specifically to cancer are actually already present in benign tissue. This has big implications for diagnosis and also potentially for deciding which bits of a cancer need treating.'
Professor Joakim Lundeberg of KTH Royal Institute of Technology, said: "Mapping thousands of tissue regions in a single experiment is an unprecedented approach to deconvolute the heterogeneity of tumours and their microenvironment. This high-resolution view impacts our way of addressing complex ecosystems such as cancer. The possibility to identify early events is particularly exciting going forward.'
Furthermore, the researchers analysed more than 150,000 regions in three prostates, two breast cancers, some skin, a lymph node and some brain tissue, and developed an algorithm to track groups of cells with similar genetic changes -clones- in their precise location. This approach enabled them to zoom right in from visible tissue through microscopic multi-cellular structures and right into the genes themselves, while keeping hold of the overall landscape of tissue.
Dr Henry Stennett, Research information manager at Cancer Research UK, who funded the research, said: 'This fascinating research challenges our understanding of how cancer develops. Using cutting-edge technology, our scientists have built an incredibly detailed 3D map of the prostate. Their results show that apparently healthy cells in the body can have the same DNA damage as cancer cells. Working out what stops them from becoming cancerous could help us to detect this disease earlier.'
The full paper, ‘Spatially resolved clonal copy number alterations in benign and malignant tissue', is published in Nature.
This research was funded by Cancer Research UK.
 Kennedy Institute of Rheumatology secures five-year major funding from the Kennedy Trust for Rheumatology Research
Kennedy Institute of Rheumatology secures five-year major funding from the Kennedy Trust for Rheumatology Research
 Oxford scientists uncover how the brain resolves emotional ambiguity
Oxford scientists uncover how the brain resolves emotional ambiguity
 Expert Comment: In Claude We Trust? Evaluating the New Constitution
Expert Comment: In Claude We Trust? Evaluating the New Constitution