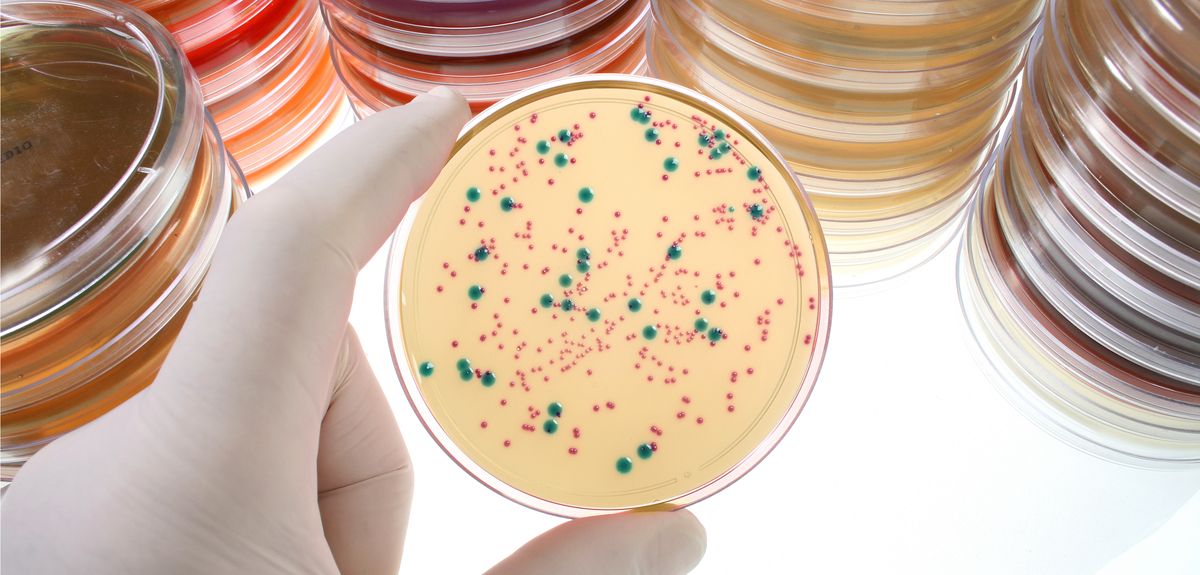
Image credit: Shutterstock
Public invited to help tackle antibiotic resistance
An online project has been launched to study antibiotic resistance in Tuberculosis (TB) with the help of the public.
The project website shows volunteers images of a series of small, circular wells, each containing M.tuberculosis (the bug that causes TB) and a different dose of an antibiotic. The users are then asked to identify in which wells the bacteria have grown, helping the researchers to determine which antibiotics are effective at killing each specific strain of TB.
Dr Philip Fowler, the lead researcher on the online BashTheBug project at the Nuffield Department of Medicine, said: 'Antibiotic resistance is a global threat, and accurately and rapidly diagnosing drug-resistant disease places a huge strain on hospital laboratories.
'Knowing which antibiotics are effective against a particular bacterial infection is crucial for effectively treating a patient, whilst also limiting the opportunity for the bug to develop antibiotic-resistance and/or to be passed onto other people.
'Cultivating and examining TB plates is a time-consuming process, but by enlisting extra help online we hope to examine over 40 million images, something we could never do on our own.'
M.tuberculosis, grows very slowly, and it can take up to two months to obtain the full diagnostic information for a patient. In March 2017, Public Health England began routinely sequencing the genomes of new suspected TB infections, and now return the same information in only 5-7 days. At the heart of this approach is a catalogue linking mutations found in the bacterial genetic code to the efficacy of different antibiotics.
The CRyPTIC project, of which BashTheBug is a part, is collecting and analysing more than 100,000 TB infection samples from across Africa, Asia, Europe and the Americas at more than a dozen centres between now and 2020. The TB genomes isolated from each sample will be sequenced and their sensitivity to a range of antibiotics tested using a specially designed culture plate, photos of which are uploaded to BashTheBug on the citizen science projects website, Zooniverse. By comparing the genetic code with the results from the culture plate for so many samples from around the world, the CRyPTIC project will build up a more complete picture of which mutations in the TB genetic code confer resistance, ultimately improving the speed and accuracy with which TB is diagnosed and treated.
TB is one of the top causes of death by infectious disease in the world, with 10.4 million cases of the disease in 2015, and 1.4 million deaths directly attributable to TB.
The CRyPTIC project is led by researchers at the NIHR Oxford Biomedical Research Centre, a partnership between the Oxford University Hospitals NHS Foundation Trust and the University of Oxford, with funding from Wellcome, The Bill & Melinda Gates Foundation and the MRC Newton Fund.
You can try out the project for yourself on the Zooniverse website, which contains instructions for how to take part.
 Oxford tops QS World University Rankings in four subjects, named overall top for Humanities
Oxford tops QS World University Rankings in four subjects, named overall top for Humanities
 Blood test may improve survival of childhood cancer in Africa
Blood test may improve survival of childhood cancer in Africa
 University of Oxford receives further £2m gift from Fondation Docteur Sadok Besrour to strengthen academic primary care in Tunisia
University of Oxford receives further £2m gift from Fondation Docteur Sadok Besrour to strengthen academic primary care in Tunisia