Enzyme structure offers new hopes for better antivirals
Publication date: 2016
The structure of a cellular enzyme that is crucial for the survival of many pathogenic viruses has been solved in a new study.
Nicole Zitzmann and members of her team, together with colleagues from Italy and France, have published their findings in PNAS (1). Their work on the enzyme, a key component of the quality control machinery that ensures that glycoproteins fold properly, opens the way for the development of potent and selective antivirals against a range of deadly diseases.
Proteins of all the major human pathogenic viruses, including Zika, dengue, influenza and Ebola, are dependent upon the host cell machinery that controls glycoproteins passing through the endoplasmic reticulum (ER). After the glycoprotein enters the ER, whether it is viral or cellular, it passes through a set of enzymes that modify its glycan chains and help it to fold properly. This is known as the calnexin cycle - calnexin, a component of the cycle, is a lectin that holds the glycoprotein so that it can fold properly. If the glycoprotein is not correctly folded on its first cycle, it must pass round the cycle again. ER α-glucosidase I and II (α-GluI and α-GluII), which sequentially remove glucose from N-linked glycans on glycoproteins, and UGGT, are the main enzymatic players in the cycle.
As Professor Zitzmann explains, the cycle has become a target for anti-virals: 'We think that viral proteins may have to go round the cycle multiple times before they are free to leave, which is much less likely to happen for normal cellular proteins. We know this is the case for HIV gp120. Our idea is to prevent viral proteins from entering the cycle by acting at α-glucosidase I and α-glucosidase II.'
Their work is currently focused on using glycomimetics called iminosugars to do this. As competitive inhibitors of α-glucosidases, these are a promising class of antivirals because they target a broad range of viruses without the risk of losing efficacy because of mutations in the viral genome. Iminosugars are currently in phase I and phase II clinical trials for dengue fever. Because iminosugars are same-site binding inhibitors, they have off-target effects and also affect intestinal α-glucosidases, which are very similar.
Whilst the focus has been on improving the efficacy of iminosugars, little work has been done on the α-GluI and α-GluII enzymes themselves with a view to developing better inhibitors. Two recent papers from the Zitzmann lab showing that inhibiting α-GluII is sufficient to inhibit dengue virus both in vitro and in vivo have shifted the focus back onto the enzymes. 'We realized that we should explore the potential of the enzymes, and in particular α-glucosidase II,' explains Professor Zitzmann. 'There are no crystal structures of any of the main mammalian enzymes of the calnexin cycle so we decided to solve the structure of α-glucosidase II, from the anti-viral angle.'
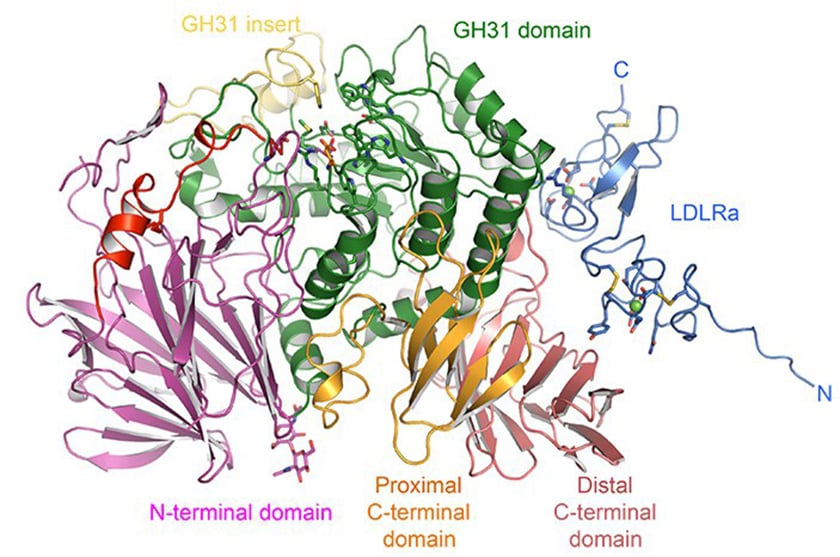
α-GluII is a heterodimer with α- and ß- subunits; the α-subunit has the active site, which binds glycan, whilst the ß-subunit retains the glycoprotein in the ER. With crystallography expertise from senior postdoc Pietro Roversi and Wellcome Trust DPhil student Alessandro Caputo, the group was able to establish the first expression system for intact mammalian α-GluII heterodimer. The mouse enzyme (Mmα-GluII) was then treated with trypsin to remove protruding loops, leading to crystals composed largely of the α-subunit and some of the ß-subunit, which retained the α/ß interface.
The group complemented this approach with small-angle X-ray scattering (SAXS), performed by part II undergraduate Souradeep Basu, which allowed them to measure the distances on the intact enzyme. By superimposing the crystal structure on the SAXS envelope, they could determine more about the enzyme's shape and detailed structure. Dominic Alonzi assayed the enzyme activity against synthetic and natural substrates.
Crystallographic studies on the mouse enzyme included Mmα-GluII in complex with its substrates as well as with antiviral iminosugar inhibitors currently in clinical trials and with a glycomimetic called 5F-glucose (synthesized by JL Kiappes from the group) that inactivates the enzyme.
Together, these studies have led to a detailed structure of α-GluII and insight into its interaction with substrates and inhibitors. When compared with the structure of an intestinal α-glucosidase, the structure has revealed some crucial differences. 'α-GluII has unique features including an 'exclusion loop' that hangs over the active site,' explains Professor Zitzmann. 'This obstructs bigger inhibitors like acarbose, which cannot fit in.'
By identifying more extensive regions that are unique to α-GluII, which iminosugars bind to - shaped by the exclusion loop and the N terminus - the group is now in a strong position to improve the current inhibitors, says Professor Zitzmann. 'Our structure gives us insight into how to design more specific and better iminosugar inhibitors. We will also be able to screen for allosteric inhibitors - chemical fragments to fit into the regions that are unique to α-GluII, which are less likely to have side-effects than same-site binding inhibitors.'
Professor Zitzmann adds that the multi-disciplinary nature of the group, comprised of crystallographers, glycobiologists and chemists, has been instrumental in its success and will be important in the future. 'Having solved the first mammalian structure of the main enzymatic players in the calnexin cycle, our target is to crystalize all three enzymes and so begin to understand how the cycle as a whole works and how we can best use this knowledge to target viruses that kill humans.'
Reference
Structures of mammalian ER α-glucosidase II capture the binding modes of broad-spectrum iminosugar antivirals. Caputo AT, Alonzi DS, Marti L, Reca IB, Kiappes JL, Struwe WB, Cross A, Basu S, Lowe ED, Darlot B, Santino A, Roversi P and Zitzmann N. PNAS July 26, 2016, doi: 10.1073/pnas.1604463113